Network measures predict neuropsychological outcome after brain injury
David Warren, Jonathan Power, Joel Bruss, Natalie Denburg, Eric Waldron, Haoxin Sun, Steve Petersen, Daniel Tranel
PNAS 2014 Sep 30; 111(39):14247-52
PubMed link
Figures (ppt)
This paper is a natural extension, and the first one we attempted, of the 2013 Neuron paper. In the 2013 Neuron paper we criticized the use of degree (and related measures) to identify important nodes in resting state data, techniques that have routinely identified portions of the default mode network as brain hubs (e.g., the precuneus). We argued that node degree may not actually index anything important in resting state correlation networks. And we argued that in resting state correlation networks, measures of signal diversity were probably indices of node importance that should roughly indicate the breadth of processes that a node participates in. So, in that paper, we highlighted several brain locations that would yield broad deficits in processing if lesioned. And our prediction was also that, based on our framework, several high-degree locations previously implicated as "brain hubs", like the precuneus or vmPFC, would not yield such broad deficits if lesioned.
So in this PNAS paper we tested that idea. We approached Daniel Tranel and his neuropsychology colleagues at the University of Iowa, where they manage the Iowa Patient Registry, a large database of patients with brain lesions lesions. This database has been growing for about 30 years, first under the Damasio's and now under Dan's leadership. The patients are imaged well after their lesion occurs and lesion maps are created by hand-marking the images. The patients also undergo a comprehensive neuropsychological assessment that includes standardized test batteries and auxiliary tests depending on the case. So we approached our colleagues at Iowa with 6 "target" locations where our 2013 Neuron paper predicted cognitive deficits would be especially broad, and together we also chose 2 "control" sites where our predictions were the opposite of the "target" locations. These control sites happen to be the precuneus and vmPFC "hubs" identified by earlier studies of node degree in resting state correlation networks. 30 patients were identified with focal lesions encompassing one of the 8 sites (their behavioral data had not yet been accessed), and the questions were:
1) would lesions to target sites yield impairment in many cognitive domains?
2) would lesions to control sites yield impairment in few cognitive impairment?
I am not a neuropsychologist, so this is where I depend on the expertise of Dan and his colleagues. Dan had 2 other blinded neuropsychologist evaluate the behavioral data of these patients, without knowing where the lesion was, and rate each subject as unimpaired, impaired, or badly impaired in each of 9 major cognitive domains (you can see the domains in the Lezak textbook Neuropsychological Assessment, which Dan also edits).
As an aside, Dan could have been one of our raters had he not been so sure early on that this project would fail - he actually unblinded himself by looking at the "target" cases once they were selected and found, to his surprise, that case after case seemed to be badly and broadly impaired. Of course he had just excluded himself from the evaluation process because he knew they were target cases. But, on the other hand, now he was definitely interested in the collaboration, and there are plenty of other clinical neuropsychologists who could do the evaluations. I digress.
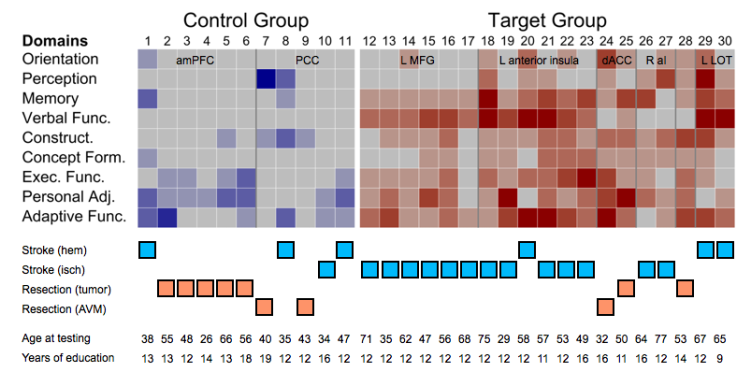
Fortunately for us, the answer was yes to both of our questions - patients with lesions at the target sites were usually impaired in about 5 of 9 cognitive domains, whereas patients with lesions to control sites were usually impaired in about 1 of 9 domains. The differences between the groups were so striking that we just showed the raw data in the paper - a matrix of patients x domains with color indicating the impairment ratings. I am told by Dan that the amount of impairment in the target cases is surprising from a neuropsychological perspective, given the lesion location and size. If our results stands as we examine more patients and sites, then perhaps what we have begun to do is to find some clinical relevance to our studies of brain network organization. When approaching a brain lesion, clinicians leverage 150 years of accumulated knowledge (e.g., the work of Luria) to set expectations about what impairments the patient will have. Perhaps there will be added value in considering the situation of the damaged tissue in terms of brain network organization.
We are expanding this study presently, seeking to replicate the findings and also extend them in new and interesting ways. David Warren, the lead author of this study and a post-doc at Iowa, was very generous to include me as a co-first author on this report.
JDP 2/1/15
Bonus materials:
There are two issues raised by this study that I would like to mention that we could only briefly discuss in the paper due to space limitations.
The first issue is whether other factors can explain the findings. In other words, should you believe the explanation we give (network properties of the absent/damaged tissue) for the dissociation between target and control lesion consequences, or could the findings be just as easily explained by other, less interesting factors?
In terms of the lesions, the control lesions were a little larger than the target regions, which would tend to work against our hypothesis. We didn't show this explicitly in the paper but there are not obvious differences between target and control lesions in terms of amount of white matter or subcortical tissue impacted (this can be a little hard to estimate since tissue can be distorted after the lesion, but it's certain that in either class of lesions there were several lesions with substantial white matter and/or subcortical gray matter involvement, and several lesions with little to no subcortical gray matter or white matter involvement). We'll quantify this more going forward as the number of lesion sites and cases investigated rises.
In terms of lesion etiology, we spent some time trying to check that the differences weren't due to some uninteresting factor. I'm placing a modified version of Figure 2 of the paper below, and combining it with some demographic information in other tables to make this point. No matter how you dice the lesions up in terms of etiology, the general dissociation between target and control lesions holds. The closest it comes to failing is when you only consider hemorrhagic stroke as a lesion etiology, where all target cases are broadly impaired, one control case is narrowly impaired, and two control cases are intermediately impaired (they are the two most impaired control cases). In all other etiologies, the cases conform well to the predictions. The Ns are low here, but this evidence doesn't suggest that etiology is what's driving our effects.
Another important factor is patient age (at the time of lesion and at the time of testing), and factors such as level of education attained. In terms of age, the control sample is younger than the target sample. But also doesn't seem to be a strong predictor of behavioral status in these data. The two worst control cases are both relatively young (38 and 35) and there patients of comparable age in the target category (32 or 29) are very broadly impaired. If one examines the older control cases (in their 50s) and comparable target cases (in their 50s), the lesions conform to predictions. So although it would have been nice if age were perfectly matched, or if the targets were younger on average than controls, it really doesn't seem that age is a primary driver of our results. Similarly, though the control patients are slightly more educated than the target patients (13.5 years vs 12 years of education), it's hard to see any influence of level of education on the findings.
Other possible explanations are that language or executive impairment could account for the disparities. The data don't strongly support these accounts either. Although most target patients do have language impairments, several do not and yet have broad impairment (no control patients have language impairment). And although ~2/3 of both control and target patients have executive impairment, the control cases with executive impairment have few other impairments, whereas the target cases uniformly have broad impairment. In the one case where a target patient (case 26) is spared in both language and executive domains, nearly every other domain is impaired.
On the whole, the single best explanation of the behavioral findings is whether the case was a target or control lesion. As our Ns increase we will be more able to document the (lack of) influence these other factors have on the behavioral findings.
The second issue that this paper raises is about interpreting brain network properties.
When I first noticed what came to be called the target locations, it was just by eye as I looked at the surface maps of the modules that had resulted from my 2011 Neuron paper. I noticed that places like the anterior insula or lateral occipito-temporal cortex had an especially large number of modules in close proximity, which got me thinking about what properties in the network would identify these possible points of convergence, what types of processing might occur there, what the consequences of lesions there would be, and whether and how such ideas could be tested. So I was all along looking for a specific property that I thought had a "cognitive" parallel. I made up one property ("system density") to summarize what I was seeing by eye, and chose another property ("participation coefficient") that should be relevant to the ideas I was developing around the target locations. And when I thought about those measures there seemed to be a straightforward link between what the measures said about the underlying signals (there are many in close proximity and/or signals here correlate strongly to multiple systems) and a story about the cognitive consequences of losing those signals.
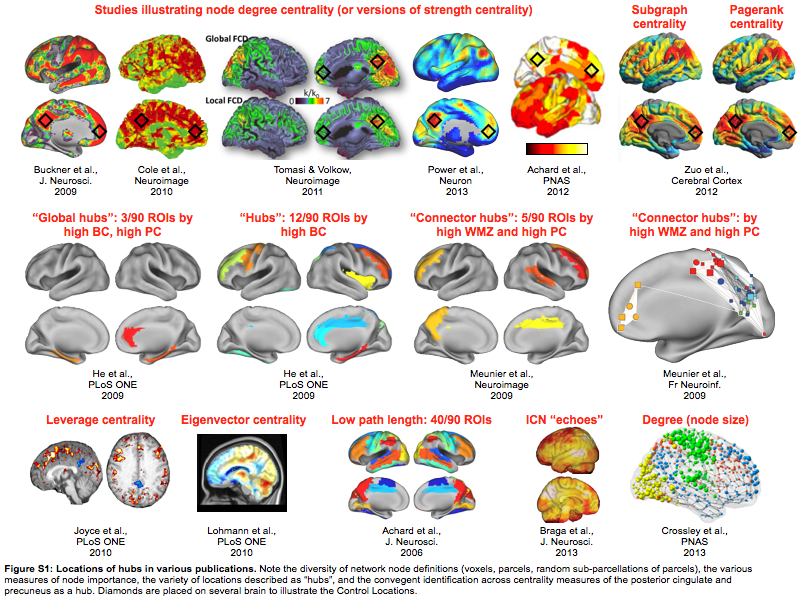
There are many network properties that can be calculated. Not all of them make sense to me in the context of correlation networks, which is what I and most other people in the resting state field use (but I've discussed that elsewhere already). The images below illustrate brain "hubs" identified in resting state data by various measures (it's a supplemental image in the PNAS paper).
There are other papers whose images could have gone here, but I think you get the idea. Pulling these images together raises points that I think are very important if you're interested in brain network organization.
First, you have to decide if you think there is a real, stable, knowable organization to the brain that is accessible by fMRI. I think there is. This means the organization should, generally, be similar across cohorts and across subjects, with wiggle room for individual variability, etc. If this organization has units, then networks should be defined so that those units are nodes. I think there are units, and I spent years trying to define the nodes as well as I knew how.
If you believe that the brain has a real organization with stable properties, then those properties should not change across studies. For example, the Meunier studies in the 2nd row of images both use the same algorithmic approach - high within module z-score and high participation coefficient - to identify "connector hubs" in resting state correlation networks. It's a little hard to tell from the images (I had to make the images, the first is reconstructed from a table and the AAL atlas, the second is a pseudoanatomical graph from the paper superimposed on a brain surface), but it doesn't really look like the same locations are being identified. If the brain has a real, stable organization, then these images cannot both be right. In this case there is a simple explanation for why the results differ - the underlying networks were defined by AAL parcels in the first case and by random subparcellation of the brain in the second case. But the point is that if the brain has real organization, then at least one of these images is inaccurate.
More generally, if you look across many studies, you'll see nearly every possible brain region highlighted as a "hub" by some study. This variety stems from (1) network definition, as was just illustrated, (2) differences in data processing, and (3) the different algorithms used to identify "hubs". Hubs can't be everywhere. By definition, they are especially important nodes.
This incoherence in the literature makes me think that two things are needed to help us move forward. First, I think we should talk less about "hubs", which is an overloaded umbrella term, and more about the specific type of node we are trying to identify with whatever measure. And second, we need to make a case for why we are considering a specific property, what it is thought to measure about brain signals or brain organization, and how we could test the implications of those claims. If the second part is not stated, how can we gain insight from these studies?
Note that the regions we used as control sites in the present study are "hubs" in many prior studies (see the diamonds in the figure). The control lesions yielded deficits in few cognitive domains. In the case of vmPFC lesions the deficits were usually emotional impairments (as expected from clinical knowledge about localization of function).
JDP 2/2/15